Our laboratory is interested in the pathogenesis of bacterial lung infections particularly tuberculosis and Legionnaires’ disease. We are examining the virulence mechanisms of bacteria using cellular, molecular and genetic techniques. Our primary research goal is to obtain a better understanding of the roles of the pathogen and host in disease. These studies should contribute to our understanding of host-pathogen interactions at the molecular and cellular level that can be used for prevention, treatment and diagnosis. We hope that through a better understanding of the mechanisms by which these organisms cause disease we can prevent some, if not all, of these infections in the future.
Tuberculosis
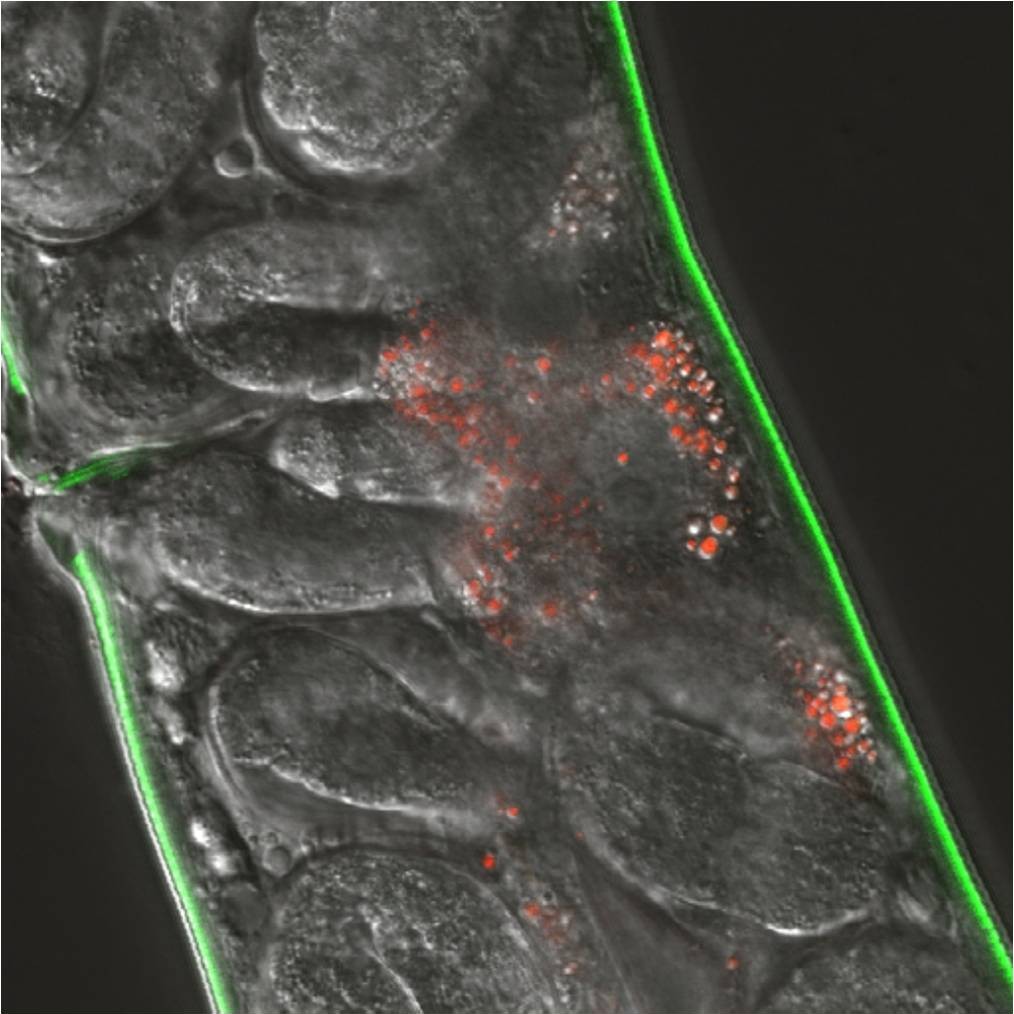
Mycobacterial research in our laboratory also focuses on the mechanisms involved in direct interactions with host cells or factors. We often use the rapid-growing mycobacteria Mycobacterium marinum as a model for tuberculosis due to its ease of use, but all studies are focused toward the important pathogenic mycobacteria, M. tuberculosis or M. avium. We have identified genetic and environmental factors that impact virulence of mycobacteria. Through the use of RICE, TnSeq and other molecular genetic strategies, we have identified many loci that are important for pathogenesis. As we develop a better understanding of the biological function of these genes we are gaining insight into the complexity of host-pathogen interactions during the different phases of disease.
Legionnaires’ Disease
Our laboratory has focused on investigation of the mechanisms of Legionella pneumophila infection of host cells and how these interactions impact their ability to cause disease. We have begun to identify both the bacterial and host genes involved in this process. By examining both sides of this interaction we hope to dissect it at the molecular and cellular levels. Clearly entry into macrophages is a complex process involving multiple bacterial and host cell components. We make mutations in both the bacterial and host genes involved to allow evaluation of each mechanism in subsequent intracellular events as well as the disease process. At present, we have identified many Legionella genes that play a role in host cell infection. The majority of these genes were identified through the use of a novel molecular approach, designated Replicating and Integrating Controlled Expression (RICE) systems. Many of the determinants isolated are involved in processing, secretion and regulation of proteins involved in entry. We have found that the Legionella strains currently used for investigation of pathogenesis differ in a number of genes that effect virulence. We have found that the differences affect adherence, entry, intracellular replication and virulence in animal models. These differences could play key roles in the sporadic epidemics caused by these bacteria.
The Host
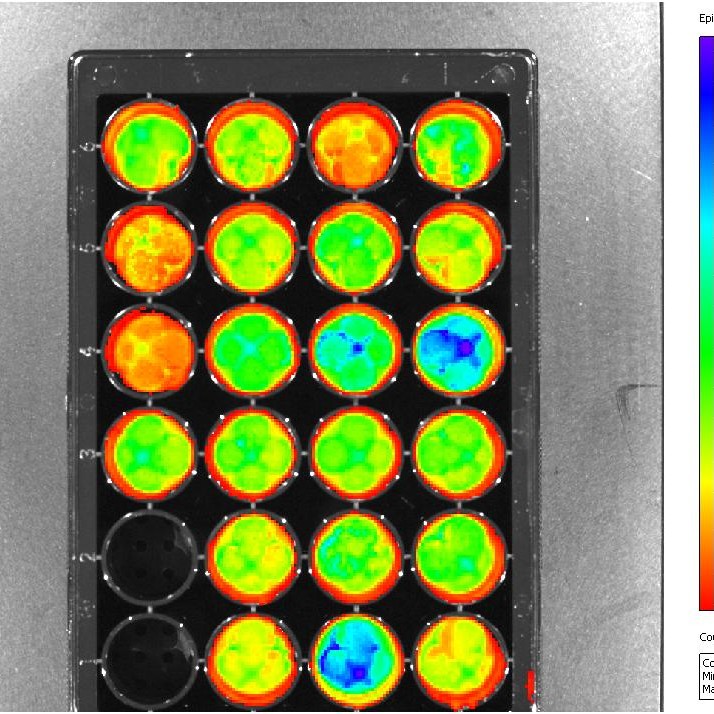
In order to better understand host factors involved in disease, we use several model systems: macrophages, the amoeba A. castellanii, mice and guinea pigs. Since amoebae are single-celled and relatively easy to grow and maintain in the laboratory, they represent a useful model system for understanding bacterial virulence mechanisms. We have identified six L. pneumophila-resistant A. castellanii (Lra) clones. Proteomic analyses allowed identification of three proteins that differ between these mutants and wild-type amoebae. In addition, we have developed a selection that allows us to isolate populations of mammalian monocytic cells that lack receptor(s) of interest. This approach combined with microarray analysis of gene expression has allowed us to evaluate the role of receptors in host cell interactions. We have identified several host factors involved in interactions with pathogens and analyzed their function in transgenic mice. Through collaborations with other groups nationally, we recently completed the guinea pig genome sequence and are using the information obtained for high-throughput sequencing and Next Generation sequencing to analyze susceptibility and resistance to infection in guinea pigs, which represent a more human-like model for disease, as compared to most other small animal models. Thus, we are taking a multi-faceted approach to the understanding of the interactions of bacteria with phagocytes from the perspective of both the bacterium and its host.